This section describes the computational facilities at the CMCF, including computer accounts, computer locations, image storage and processing capabilities. We also describe data transfer options, image viewing options and include a list of helpful Linux commands.
To see details about our software, please visit:
- MxDC - CMCF data collection software
- MxLIVE - CMCF sample and experiment tracking software
- Autoprocess - CMCF automated data processing pipeline
- Crystallography Software - external software that Users have access to.
Computational Facilities
User Accounts
New user groups to the CMCF will be issued an account used to log in to:
- beamline user computers,
- NX client for remote control, and
- MxLIVE on the web.
Any images generated on the beamline will be saved under the logged-in user's account and will only be visible to that user.
Computer Locations
There are two user workstations available in the CMCF-ID Control Room and two workstations in the CMCF-BM Control Room (see the map for locations) Additional computers are also available in the following locations:
- Experimental Hutch - This may be used to centre samples before locking up the hutch, but should not be used to collect data.
- Sample Prep Room - This should not be used for data collection.
Workstations marked "Staff" should only be used by beamline staff.
Data hierarchy, storage, and retention
As data is collected on a CMCF beamline, images are saved and stored under the user's home directory ( /users/<user_name>/...) and organized by session (...<session_name>/<group_name>/<sample_name>/..) Data for individual crystals is grouped by type (eg. /data and /screen for image files, /proc-native and /proc-screen for autoprocess results). Default session_name, group_name, and sample_name values are specified based on shipment details in MxLive. User directories and data are only visible to that individual user account.
Data stored at CMCF will be accessible in the user's account for 1 year based on the session date, or until space is needed. When space is needed, old data will be removed on a first-in, first-out policy. While current resources allow for 2+ years of storage, this is not guaranteed and users are encouraged to backup their data soon after their session to avoid any loss.
Data Transfer Options
MxLIVE
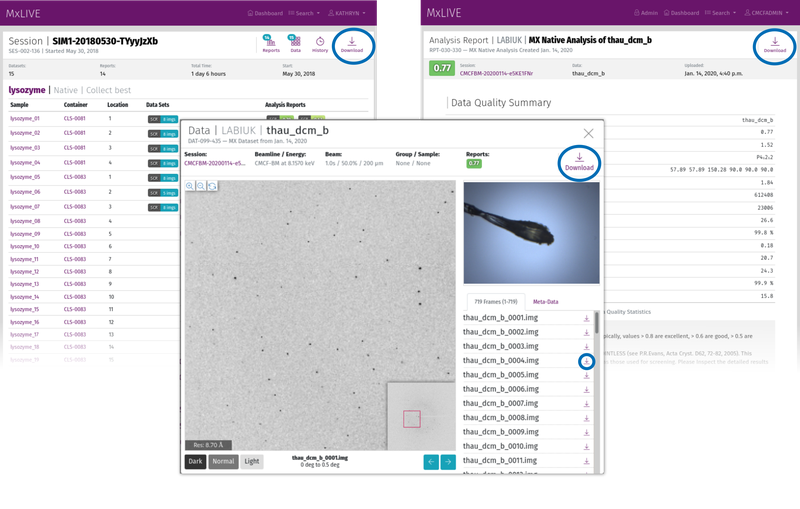
The most convenient method of data transfer is using MxLIVE. Login with your beamline user account and download individual datasets, reports (including all processing files), or your complete session directory. Find the "Download" icon at the top-right hand corner when viewing your dataset, report, or session, or download individual diffraction frames from a dataset by finding the download icon on the right-hand side of each frame in the frame list.
An archive of the entire dataset, report, or session will be created and streamed to you through your browser. Note that session downloads include all datasets and reports from that session, as long as they are available in our on-site data storage. Note: Manual data processing does not appear in MxLive but will be downloaded alongside the rest of your session.
Hard Drive
Data, including image files and processing files, can be transferred to a user-supplied USB external hard drive and shipped along with your samples. An archiver script has been set up to automate the task of copying data. Set up a directory in which all of your data will be collected. Connect your drive to a User Workstation (we recommend a different workstation than the one you are collecting data on). From the User Workstation your drive is connected to, open a local terminal (not a data processing terminal), and simply type
archiver /user/[user_name]/[session_folder]/ /run/media/[user_name]/[drivename]/[session_folder]
The archiver script will then automatically backup all of the data collected in subsequent subdirectories as you collect.
NOTE:
• Any USB drive should be NTFS or FAT32. EXT2-4 drives are supported provided the permissions on the target directory have been previously set to be world-writable. This can be done with the following command using administrator privilegeschmod 777 <target_directory>• A 3600 image dataset and processing result is compressed to ~4-6 GB. Please ensure your drive has sufficient space for your entire session.• FAT32 hard drives formatted using Windows XP (or before) are limited to partitions below 32, 768 MB and are not appropriate for use with .h5 files. Please reformat your drive using an x64 system (note: reformatting drives will delete all data currently stored on the disk).
• Firewire is not supported at this time.
Data Transfer via SSH
Users who have uploaded their public ssh-key to their profile on MxLIVE can also request temporary access to the data transfer server by emailing their support contact. Data transfer can then be completed using either of the following commands. SSH keys are accepted in RSA or DSA format and should be of the format ssh-rsa <your_public_key_string> , omitting any other information present in the PubKey (.pub) file. SSH key-pairs can be generated using ssh-keygen -t rsa -b 4096 .
scp -r -C -P [port] [user_name]@cmcf.lightsource.ca:/[absolutepath]/[to]/[session_directory]/* /[location]/[on]/[localmachine]
(Ideal for only a few files.)
rsync -avzh -e 'ssh -p [port]' [user_name]@cmcf.lightsource.ca:/[absolutepath]/[to]/[session_directory]/* /[location]/[on]/[localmachine]
(Ideal for large downloads and/or unstable connections. This command can be repeated as new data is generated. This command is not available in Windows by default)
Double check that you have enough disk space and have correctly entered the file path prior to running rsync or scp. Files with the same name can be easily overwritten if incorrectly executed.
Viewing Diffraction Images
Viewing Images at CMCF
- To view diffraction images outside of the MXDC data collection software, you may use the icon to launch the Diffraction Frame Viewer
- Or simply type
imgview [filename] - To view the header information stored in an image file, type
imginfo [fileneame] - Image viewing is also supported using
imosflm
Viewing Images on Your PC
Diffraction images collected at CMCF are generated as single frame .cbf (for CMCF-BM) or multi-frame .h5 (for CMCF-ID) files. Viewing of images is generally supported in up-to-date versions of MosFlm and HKL2000/3000, provided the data metadata master file is also included. HDF5 files contain image data in a hierarchical format, and will often require a separate viewing program to interpret. More details on the HDF5 format can be found here.
Linux Basics
Note: Most commands in Linux are case-sensitive.
Execute command but pause after each screenful of output
command |morecommand |less
Launch command in the background
(useful for GUI programs such as gedit)
command &
Alternative way of running programs in the background
Press CTRL-Z after running command and then type bg
Launch an easy-to-use text-based file editor
nano filename
[If you know how to use Vi or Emacs, why are you reading this page?]
Display the contents of filename
cat filename
Display the last ten lines of filename
tail filename
Reset the current terminal
(useful if you cat a binary file by mistake and the terminal gets scambled)
reset
List all files in the current directory, sorting by date
ls -lrt
List all files in the current directory, sorting by size
ls -lrS
Search the current directory and all subdirectories for filename
find . -name filename
See the free space remaining on all mounted partitions
df -h
To see the size of directories
du -sh *